Most importantly, download the Docker images in advance, the conference venue's Wi-Fi may be slow and unreliable. Looking forward to meeting you — Mitchell Shiell, Ontario Institute for Cancer Research, mshiell@oicr.on.ca
IBC Workshop Prerequisites
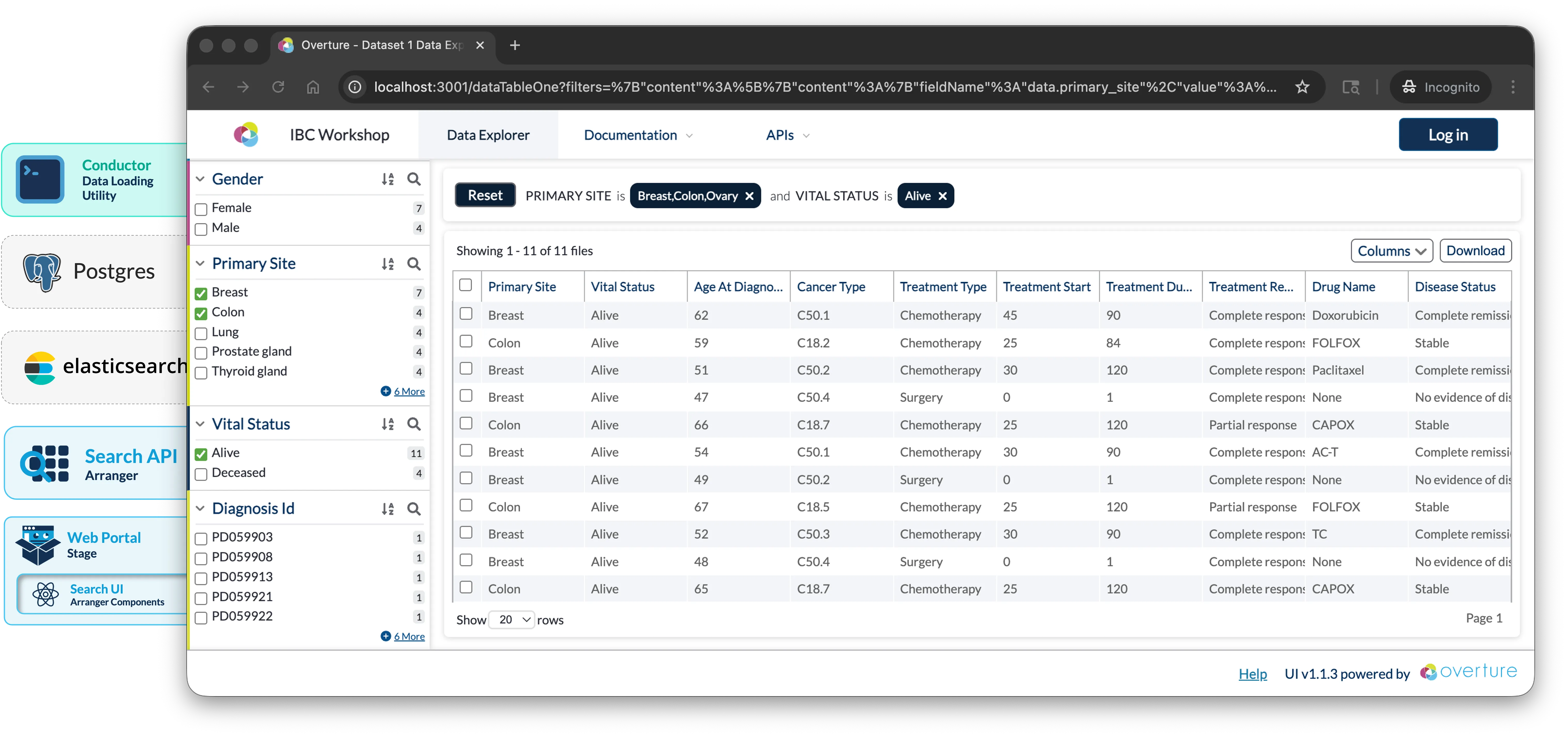
This workshop has been developed as part of the 19th Annual International Biocuration Conference, it will guide you through building a foundational data discovery portal for tabular CSV data using Elasticsearch, Arranger, and Stage.
If you're attending, feel free to drop a quick introduction before the day, this helps tailor the session to the room. Entirely optional.
Objectives:
- Deploy a functional data discovery portal using Elasticsearch, GraphQL, Arranger, and Stage
- Configure search interfaces and indices tailored to tabular datasets
- Gain familiarity with the tools needed to adapt this portal to your own data
- Understand deployment options for making portals accessible on institutional networks and beyond
Prerequisites
The following software should be installed and verified before the workshop:
1. Git git --version returns a version number
Download from git-scm.com if the command is not recognised.
2. Docker Desktop (28.0.0 or later)
- macOS / Windows: Download from docker.com/products/docker-desktop
- Linux: Follow the Docker Engine install guide
Once installed, open Docker Desktop → Settings → Resources and set:
- CPUs: 4+ cores (8 recommended)
- Memory: 8 GB minimum
- Disk: 10 GB+ available
Please ensure docker --version and docker compose version both return version numbers, and Docker Desktop is running with 4+ CPUs and 8 GB+ memory allocated
3. Docker images pre-downloaded Most time-consuming step, run these before the workshop
Pull the required Docker images now to avoid slow downloads during the workshop:
docker pull alpine/curl:8.8.0
docker pull postgres:15-alpine
docker pull docker.elastic.co/elasticsearch/elasticsearch:7.17.27
docker pull ghcr.io/overture-stack/arranger-server:4919f736
docker pull ghcr.io/overture-stack/conductor:171d9ce
docker pull node:18-alpine
Verify all six downloaded:
docker images | grep -E "alpine/curl|postgres|elasticsearch|arranger-server|conductor|node"
You should see all six images listed.
4. Repository cloned: git clone -b IBCworkshop https://github.com/overture-stack/prelude.git
The prelude repository contains everything needed for this workshop: Docker Compose configuration, the Conductor wrapper script, and sample data. Clone it once before the workshop and you won't need internet access for the hands-on portion.
git clone -b IBCworkshop https://github.com/overture-stack/prelude.git
5. (Windows only) WSL2 configured with Docker Desktop integration enabled
- Install WSL2
- Use Ubuntu or another Linux distribution within WSL2
- Enable Docker Desktop's WSL2 integration (Docker Desktop → Settings → Resources → WSL Integration)
- Run all workshop commands from a Bash terminal inside WSL2, not PowerShell or Command Prompt. To open one, search for your Linux distribution (e.g. "Ubuntu") in the Start menu.
Optional Prerequisites
These are not required but will make the workshop easier to follow:
6. (Optional) Elasticvue: browser-based Elasticsearch GUI
Elasticvue is a browser-based Elasticsearch GUI useful for inspecting indices, browsing documents, and troubleshooting. It is not required but helpful for understanding what's happening inside Elasticsearch during the workshop.
Install it as a browser extension or standalone app.
7. (Optional) PostgreSQL GUI client
8. (Optional) Bring your own data: CSV file
If you have a tabular dataset you'd like to use during or after the workshop, bring it as a CSV file. During the workshop we will use demo data, but the final section covers adapting the portal to your own dataset.
Schedule
Venue: Pacific 2
| Time | Section | Description |
|---|---|---|
| 2:00–2:20 | Introduction & Overview | Workshop objectives, run the pre-built demo, and architecture walkthrough |
| 2:20–3:30 | Building Your Portal | Prepare data, generate configurations, wire up Docker, Launch & Load data |
| 3:30–3:40 | Break | Stretch break |
| 3:40–4:00 | Wrap-Up | Customize the portal, discuss next steps, and Q&A |
Support
| During the workshop | You can use this slack link for additional support |
| Before or after | community support channels or contact@overture.bio |
| Bug reports | GitHub Issues |
Facilitator: Mitchell Shiell, Ontario Institute for Cancer Research, mshiell@oicr.on.ca
Verification Checklist
Before the workshop, confirm:
git --versionreturns a version numberdocker --versionreturns 28.0.0 or laterdocker compose versionreturns a version number- Docker Desktop is running with 4+ CPUs and 8 GB+ memory allocated
- All six Docker images are downloaded (
docker images) — this is the most time-consuming step, do it before the day - The repository is cloned and you can
cdinto it - (Windows only) WSL2 is configured and Docker integration is enabled
Troubleshooting: If you run into issues before the workshop, reach out via the community support channels or email mshiell@oicr.on.ca.